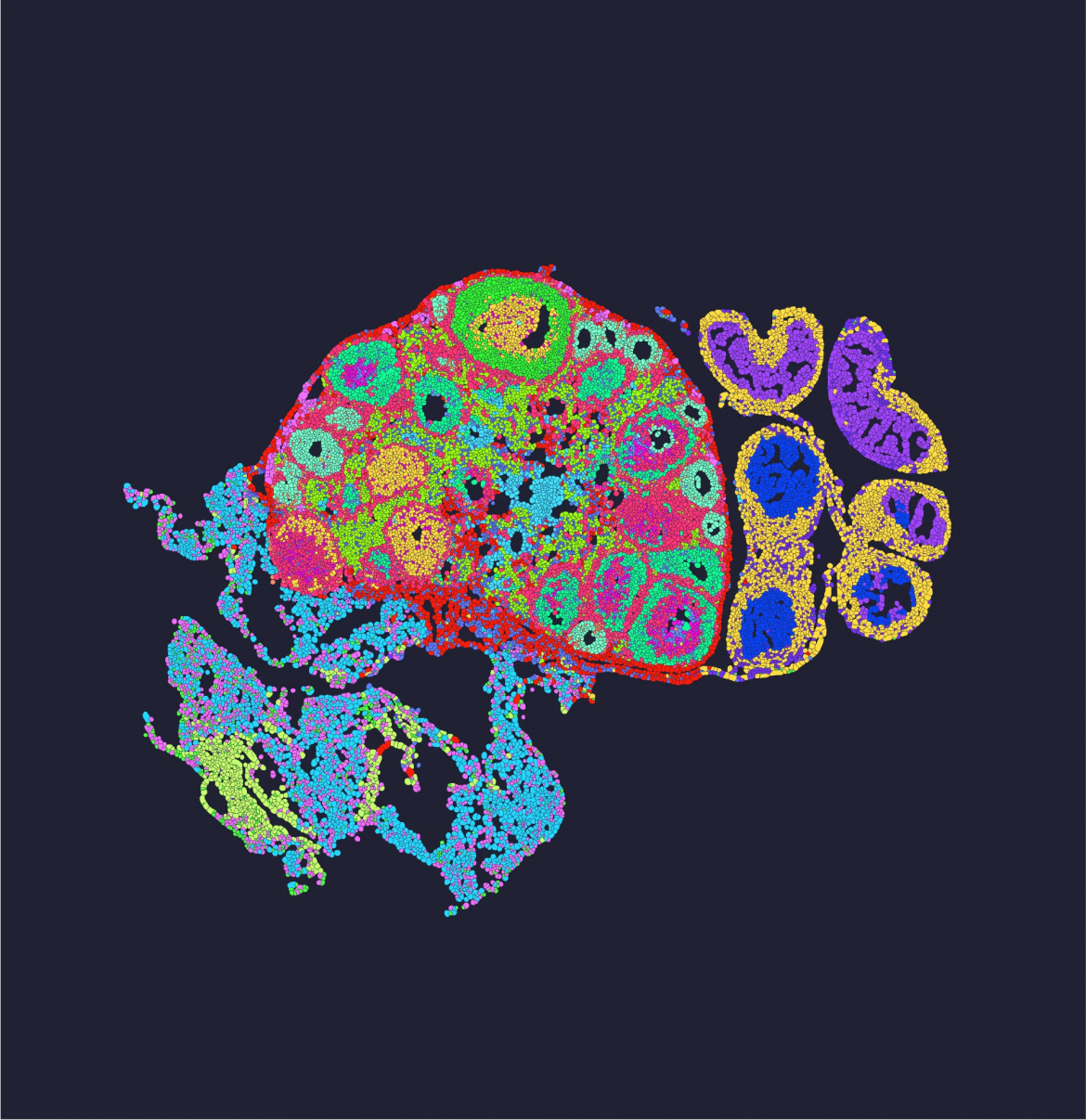
Examining gene activity through transcriptomics has been a cornerstone of biological research. Traditionally, bulk RNA-seq has served as the primary method. However, a newer approach, spatial omics transcriptomics, provides a different perspective. Platforms like STOmics have advanced this field by offering tools that capture data with spatial context, moving beyond the limitations of older techniques. This comparison outlines the fundamental distinctions between these two transcriptomics solution pathways.
The Dimension of Data Collection
A primary distinction lies in data granularity. Bulk RNA-seq requires homogenizing a tissue sample, blending all cell types together. The resulting data is an average expression profile, losing all information about where specific signals originated. In contrast, spatial omics transcriptomics platforms analyze intact tissue sections. STOmics’ Stereo-seq technology, for example, allows measurement across a complete tissue slice, preserving the native architecture and generating maps of gene expression.
Contextual Information and Tissue Architecture
The loss of spatial context in bulk RNA-seq can obscure critical biological insights. It cannot determine if a gene is active in a specific cell layer, a tumor core, or an immune infiltrate. A spatial omics transcriptomics approach directly addresses this. It links transcriptomic data to precise coordinates, enabling researchers to see how gene expression patterns relate to tissue morphology and cellular neighborhoods, which is vital for understanding complex diseases and development.
Applications and Research Outcomes
These methodological differences guide their applications. Bulk RNA-seq excels at profiling overall expression changes between large sample groups, such as diseased versus healthy organs. A comprehensive spatial omics transcriptomics workflow, however, is designed for questions where location is paramount. It is used to discover novel cell interactions, trace cell lineages in situ, and characterize the tumor microenvironment, providing a spatially resolved transcriptomics solution that bulk methods cannot offer.
For researchers requiring a complete view of tissue function, the shift from a blended average to a spatially aware map represents a significant evolution. STOmics provides the integrated tools—from sequencing to analysis—that enable this detailed, location-based investigation into the transcriptome, distinguishing it from conventional analytical methods.